Installation
Install and load rnoaa into the R session. Stable version from CRAN
install.packages("rnoaa")
Or development version from Github:
install.packages("devtools")
devtools::install_github("ropensci/rnoaa")
library('plyr')
library('rnoaa')
Usage
National Climatic Data Center (NCDC) data
Get info on a station by specifying a datasetid, locationid, and stationid
ncdc_stations(datasetid='GHCND', locationid='FIPS:12017', stationid='GHCND:USC00084289')
#> $meta
#> NULL
#>
#> $data
#> elevation mindate maxdate latitude name
#> 1 12.2 1899-02-01 2015-07-06 28.8029 INVERNESS 3 SE, FL US
#> datacoverage id elevationUnit longitude
#> 1 1 GHCND:USC00084289 METERS -82.3126
#>
#> attr(,"class")
#> [1] "ncdc_stations"
Search for data and get a data.frame
out <- ncdc(datasetid='NORMAL_DLY', datatypeid='dly-tmax-normal', startdate = '2010-05-01', enddate = '2010-05-10')
See a data.frame
out$data
#> Source: local data frame [25 x 5]
#>
#> date datatype station value fl_c
#> 1 2010-05-01T00:00:00 DLY-TMAX-NORMAL GHCND:AQW00061705 869 C
#> 2 2010-05-01T00:00:00 DLY-TMAX-NORMAL GHCND:CAW00064757 607 Q
#> 3 2010-05-01T00:00:00 DLY-TMAX-NORMAL GHCND:CQC00914080 840 R
#> 4 2010-05-01T00:00:00 DLY-TMAX-NORMAL GHCND:CQC00914801 858 R
#> 5 2010-05-01T00:00:00 DLY-TMAX-NORMAL GHCND:FMC00914395 876 P
#> 6 2010-05-01T00:00:00 DLY-TMAX-NORMAL GHCND:FMC00914419 885 P
#> 7 2010-05-01T00:00:00 DLY-TMAX-NORMAL GHCND:FMC00914446 885 P
#> 8 2010-05-01T00:00:00 DLY-TMAX-NORMAL GHCND:FMC00914482 868 R
#> 9 2010-05-01T00:00:00 DLY-TMAX-NORMAL GHCND:FMC00914720 899 R
#> 10 2010-05-01T00:00:00 DLY-TMAX-NORMAL GHCND:FMC00914761 897 P
#> .. ... ... ... ... ...
Plot data, super simple, but it's a start
out <- ncdc(datasetid='NORMAL_DLY', stationid='GHCND:USW00014895', datatypeid='dly-tmax-normal', startdate = '2010-01-01', enddate = '2010-12-10', limit = 300)
ncdc_plot(out)
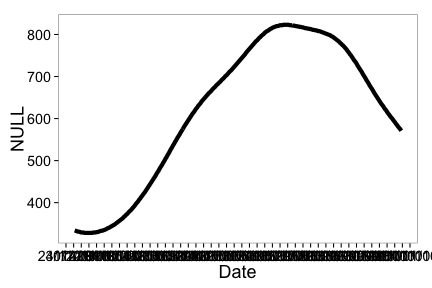
Note that the x-axis tick text is not readable, but see futher down in tutorial for how to adjust that.
More on plotting
Example 1
Search for data first, then plot
out <- ncdc(datasetid='GHCND', stationid='GHCND:USW00014895', datatypeid='PRCP', startdate = '2010-05-01', enddate = '2010-10-31', limit=500)
Default plot
ncdc_plot(out)
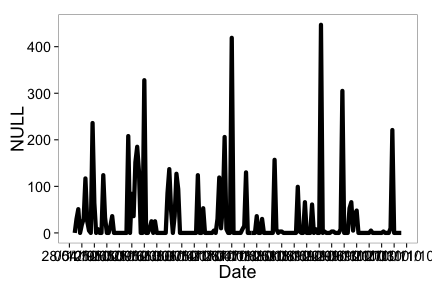
Create 14 day breaks
ncdc_plot(out, breaks="14 days")
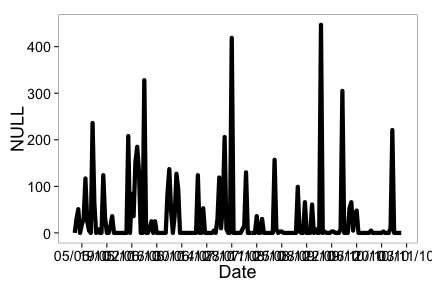
One month breaks
ncdc_plot(out, breaks="1 month", dateformat="%d/%m")
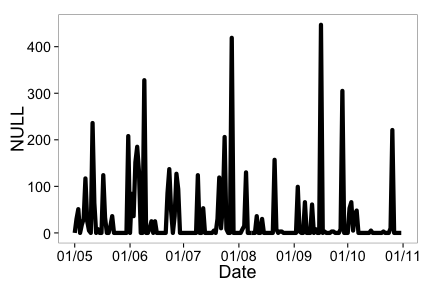
Example 2
Search for data
out2 <- ncdc(datasetid='GHCND', stationid='GHCND:USW00014895', datatypeid='PRCP', startdate = '2010-05-01', enddate = '2010-05-03', limit=100)
Make a plot, with 6 hour breaks, and date format with only hour
ncdc_plot(out2, breaks="6 hours", dateformat="%H")
Combine many calls to noaa function
Search for two sets of data
out1 <- ncdc(datasetid='GHCND', stationid='GHCND:USW00014895', datatypeid='PRCP', startdate = '2010-03-01', enddate = '2010-05-31', limit=500)
out2 <- ncdc(datasetid='GHCND', stationid='GHCND:USW00014895', datatypeid='PRCP', startdate = '2010-09-01', enddate = '2010-10-31', limit=500)
Then combine with a call to ncdc_combine
df <- ncdc_combine(out1, out2)
head(df[[1]]); tail(df[[1]])
#> Source: local data frame [6 x 8]
#>
#> date datatype station value fl_m fl_q fl_so
#> 1 2010-03-01T00:00:00 PRCP GHCND:USW00014895 0 T 0
#> 2 2010-03-02T00:00:00 PRCP GHCND:USW00014895 0 T 0
#> 3 2010-03-03T00:00:00 PRCP GHCND:USW00014895 0 T 0
#> 4 2010-03-04T00:00:00 PRCP GHCND:USW00014895 0 0
#> 5 2010-03-05T00:00:00 PRCP GHCND:USW00014895 0 0
#> 6 2010-03-06T00:00:00 PRCP GHCND:USW00014895 0 0
#> Variables not shown: fl_t (chr)
#> Source: local data frame [6 x 8]
#>
#> date datatype station value fl_m fl_q fl_so
#> 1 2010-10-26T00:00:00 PRCP GHCND:USW00014895 221 0
#> 2 2010-10-27T00:00:00 PRCP GHCND:USW00014895 0 0
#> 3 2010-10-28T00:00:00 PRCP GHCND:USW00014895 0 T 0
#> 4 2010-10-29T00:00:00 PRCP GHCND:USW00014895 0 T 0
#> 5 2010-10-30T00:00:00 PRCP GHCND:USW00014895 0 0
#> 6 2010-10-31T00:00:00 PRCP GHCND:USW00014895 0 0
#> Variables not shown: fl_t (chr)
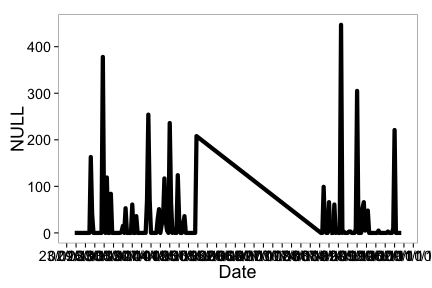
Then plot - the default passing in the combined plot plots the data together. In this case it looks kind of weird since a straight line combines two distant dates.
ncdc_plot(df)
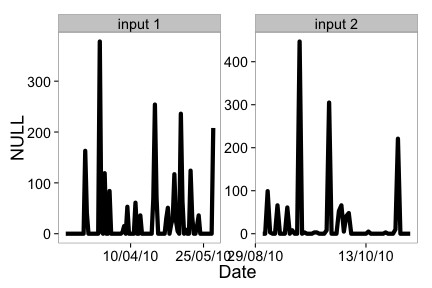
But we can pass in each separately, which uses facet_wrap in ggplot2 to plot each set of data in its own panel.
ncdc_plot(out1, out2, breaks="45 days")
ERDDAP data
ERDDAP data is now avialable through the
rerddappackage
Severe Weather Data Inventory (SWDI) data
Search for nx3tvs data from 5 May 2006 to 6 May 2006
swdi(dataset='nx3tvs', startdate='20060505', enddate='20060506')
#> $meta
#> $meta$totalCount
#> numeric(0)
#>
#> $meta$totalTimeInSeconds
#> [1] 0.022
#>
#>
#> $data
#> Source: local data frame [25 x 8]
#>
#> ztime wsr_id cell_id cell_type range azimuth max_shear
#> 1 2006-05-05T00:05:50Z KBMX Q0 TVS 7 217 403
#> 2 2006-05-05T00:10:02Z KBMX Q0 TVS 5 208 421
#> 3 2006-05-05T00:12:34Z KSJT P2 TVS 49 106 17
#> 4 2006-05-05T00:17:31Z KSJT B4 TVS 40 297 25
#> 5 2006-05-05T00:29:13Z KMAF H4 TVS 53 333 34
#> 6 2006-05-05T00:31:25Z KLBB N0 TVS 51 241 24
#> 7 2006-05-05T00:33:25Z KMAF H4 TVS 52 334 46
#> 8 2006-05-05T00:37:37Z KMAF H4 TVS 50 334 34
#> 9 2006-05-05T00:41:51Z KMAF H4 TVS 51 335 29
#> 10 2006-05-05T00:44:33Z KLBB N0 TVS 46 245 35
#> .. ... ... ... ... ... ... ...
#> Variables not shown: mxdv (chr)
#>
#> $shape
#> shape
#> 1 POINT (-86.8535716274277 33.0786326913943)
#> 2 POINT (-86.8165772540846 33.0982820681588)
#> 3 POINT (-99.5771091971025 31.1421609654838)
#> 4 POINT (-101.188161700093 31.672392833416)
#> 5 POINT (-102.664426480293 32.7306917937698)
#> 6 POINT (-102.70047613441 33.2380072329615)
#> 7 POINT (-102.6393683028 32.7226656893341)
#> 8 POINT (-102.621904684258 32.6927081076156)
#> 9 POINT (-102.614794815627 32.714139844846)
#> 10 POINT (-102.643380529494 33.3266446067682)
#> 11 POINT (-102.597961935071 32.6839260102062)
#> 12 POINT (-102.613894688178 33.3526192273658)
#> 13 POINT (-102.567153417051 32.6956373348052)
#> 14 POINT (-102.551596970251 32.664939580306)
#> 15 POINT (-102.586119971014 33.4287323151248)
#> 16 POINT (-102.499638479193 32.6644438090742)
#> 17 POINT (-102.5485490063 33.4398330734778)
#> 18 POINT (-102.51446954228 32.6597119240996)
#> 19 POINT (-102.559031583693 32.7166090376869)
#> 20 POINT (-102.492174522228 33.4564626989719)
#> 21 POINT (-102.463540844324 32.6573739036181)
#> 22 POINT (-102.510349454162 33.5066366303981)
#> 23 POINT (-102.448763863447 32.6613484943994)
#> 24 POINT (-102.42842159557 32.649061124799)
#> 25 POINT (-102.504158884526 32.7162751126854)
#>
#> attr(,"class")
#> [1] "swdi"
Use an id
out <- swdi(dataset='warn', startdate='20060506', enddate='20060507', id=533623)
list(out$meta, head(out$data), head(out$shape))
#> [[1]]
#> [[1]]$totalCount
#> numeric(0)
#>
#> [[1]]$totalTimeInSeconds
#> [1] 0.171
#>
#>
#> [[2]]
#> Source: local data frame [6 x 6]
#>
#> ztime_start ztime_end id warningtype
#> 1 2006-05-05T22:53:00Z 2006-05-06T00:00:00Z 397428 SEVERE THUNDERSTORM
#> 2 2006-05-05T22:55:00Z 2006-05-06T00:00:00Z 397429 SEVERE THUNDERSTORM
#> 3 2006-05-05T22:55:00Z 2006-05-06T00:00:00Z 397430 SEVERE THUNDERSTORM
#> 4 2006-05-05T22:57:00Z 2006-05-06T00:00:00Z 397431 SEVERE THUNDERSTORM
#> 5 2006-05-05T23:03:00Z 2006-05-06T00:00:00Z 397434 SEVERE THUNDERSTORM
#> 6 2006-05-05T23:14:00Z 2006-05-06T00:15:00Z 397437 SEVERE THUNDERSTORM
#> Variables not shown: issuewfo (chr), messageid (chr)
#>
#> [[3]]
#> shape
#> 1 POLYGON ((-93.27 30.38, -93.29 30.18, -93.02 30.18, -93.04 30.37, -93.27 30.38))
#> 2 POLYGON ((-101.93 34.74, -101.96 34.35, -101.48 34.42, -101.49 34.74, -101.93 34.74))
#> 3 POLYGON ((-100.36 33.03, -99.99 33.3, -99.99 33.39, -100.28 33.39, -100.5 33.18, -100.51 33.02, -100.45 32.97, -100.37 33.03, -100.36 33.03))
#> 4 POLYGON ((-102.8 30.74, -102.78 30.57, -102.15 30.61, -102.15 30.66, -101.92 30.68, -102.07 30.83, -102.8 30.74))
#> 5 POLYGON ((-103.02 32.94, -103.03 32.66, -102.21 32.53, -102.22 32.95, -103.02 32.94))
#> 6 POLYGON ((-101.6 33.32, -101.57 33.31, -101.57 33.51, -101.65 33.51, -101.66 33.5, -101.75 33.5, -101.77 33.49, -101.84 33.49, -101.84 33.32, -101.6 33.32))
Get all 'plsr' within the bounding box (-91,30,-90,31)
swdi(dataset='plsr', startdate='20060505', enddate='20060510', bbox=c(-91,30,-90,31))
#> $meta
#> $meta$totalCount
#> numeric(0)
#>
#> $meta$totalTimeInSeconds
#> [1] 0
#>
#>
#> $data
#> Source: local data frame [5 x 8]
#>
#> ztime id event magnitude city
#> 1 2006-05-09T02:20:00Z 427540 HAIL 1 5 E KENTWOOD
#> 2 2006-05-09T02:40:00Z 427536 HAIL 1 MOUNT HERMAN
#> 3 2006-05-09T02:40:00Z 427537 TSTM WND DMG -9999 MOUNT HERMAN
#> 4 2006-05-09T03:00:00Z 427199 HAIL 0 FRANKLINTON
#> 5 2006-05-09T03:17:00Z 427200 TORNADO -9999 5 S FRANKLINTON
#> Variables not shown: county (chr), state (chr), source (chr)
#>
#> $shape
#> shape
#> 1 POINT (-90.43 30.93)
#> 2 POINT (-90.3 30.96)
#> 3 POINT (-90.3 30.96)
#> 4 POINT (-90.14 30.85)
#> 5 POINT (-90.14 30.78)
#>
#> attr(,"class")
#> [1] "swdi"
Sea ice data
Map all years for April only for North pole
urls <- seaiceeurls(mo='Apr', pole='N')[1:10]
out <- lapply(urls, seaice)
names(out) <- seq(1979,1988,1)
df <- ldply(out)
library('ggplot2')
ggplot(df, aes(long, lat, group=group)) +
geom_polygon(fill="steelblue") +
theme_ice() +
facet_wrap(~ .id)
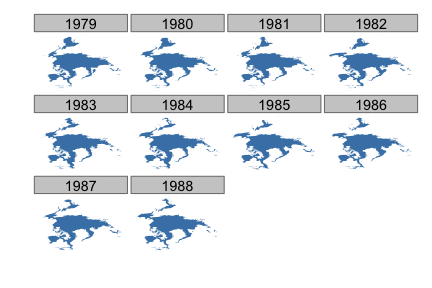
IBTrACS storm data
Get NOAA wind storm tabular data, metadata, or shp files from International Best Track Archive for Climate Stewardship (IBTrACS). See http://www.ncdc.noaa.gov/ibtracs/index.php?name=numbering for more.
Metadata
There are two datasets stored in the package. By default storm_meta() gives metadata describing columns of the datasets returned.
head( storm_meta() )
#> Column_number Column_name units Shapefile_pt_flag
#> 1 1 Serial_Num N/A 1
#> 2 2 Season Year 1
#> 3 3 Num # 1
#> 4 4 Basin BB 1
#> 5 5 Sub_basin BB 1
#> 6 6 Name N/A 1
#> Shapefile_pt_attribute_name shapefile_att_type shapefile_att_len
#> 1 Serial_Num 7 13
#> 2 Season 3 4
#> 3 Num 3 5
#> 4 Basin 7 3
#> 5 Sub_basin 7 3
#> 6 Name 7 57
#> shapefile_att_prc
#> 1 0
#> 2 0
#> 3 0
#> 4 0
#> 5 0
#> 6 0
Or you can get back a dataset of storm names, including storm ids and their names.
head( storm_meta("storm_names") )
#> id name
#> 1 1842298N11080 NOT NAMED(td9636)
#> 2 1845336N10074 NOT NAMED(td9636)
#> 3 1848011S09079 NOT NAMED(td9636)
#> 4 1848011S09080 XXXX848003(reunion)
#> 5 1848011S15057 XXXX848002(reunion)
#> 6 1848011S16057 NOT NAMED(td9636)
Tabular data
You can get tabular data for basins, storms, or years, (or all data). storm_data() and the next function storm_shp() figure out what files to get, and gets them from an ftp server, and saves them to your machine. Do let us know if you have any problems with paths on your machine, and we'll fix 'em. The result from storm_data() is a dplyr-like data.frame with a easy summary that makes large datasets easy to view.
First, by basin (one of EP, NA, NI, SA, SI, SP, or WP)
storm_data(year=1941)
#> <path>~/.rnoaa/storms/year/Year.1941.ibtracs_all.v03r06.csv
#>
#> <NOAA Storm Data>
#> Size: 1766 X 195
#>
#> serial_num season num basin sub_basin name iso_time nature latitude
#> 1 1940215S18149 1941 1 SP EA NOT NAMED 1940-08-01 12:00:00 NR -999
#> 2 1940215S18149 1941 1 SP EA NOT NAMED 1940-08-01 18:00:00 NR -999
#> 3 1940215S18149 1941 1 SP EA NOT NAMED 1940-08-02 00:00:00 NR -999
#> 4 1940215S18149 1941 1 SP EA NOT NAMED 1940-08-02 06:00:00 NR -999
#> 5 1940215S18149 1941 1 SP EA NOT NAMED 1940-08-02 12:00:00 NR -999
#> 6 1940215S18149 1941 1 SP EA NOT NAMED 1940-08-02 18:00:00 NR -999
#> 7 1940215S18149 1941 1 SP EA NOT NAMED 1940-08-03 00:00:00 NR -999
#> 8 1940215S18149 1941 1 SP EA NOT NAMED 1940-08-03 06:00:00 NR -999
#> 9 1940215S18149 1941 1 SP EA NOT NAMED 1940-08-03 12:00:00 NR -999
#> 10 1940215S18149 1941 1 SP EA NOT NAMED 1940-08-03 18:00:00 NR -999
#> .. ... ... ... ... ... ... ... ... ...
#> Variables not shown: longitude (dbl), wind.wmo. (dbl), pres.wmo. (dbl), center (chr),
#> wind.wmo..percentile (dbl), pres.wmo..percentile (dbl), track_type (chr),
#> latitude_for_mapping (dbl), longitude_for_mapping (dbl), current.basin (chr), hurdat_atl_lat
#> (dbl), hurdat_atl_lon (dbl), hurdat_atl_grade (dbl), hurdat_atl_wind (dbl), hurdat_atl_pres
#> (dbl), td9636_lat (dbl), td9636_lon (dbl), td9636_grade (dbl), td9636_wind (dbl),
Buoy data
Find out what buoys are available in a dataset
head(buoys(dataset = "cwind"))
#> id
#> 1 41001
#> 2 41002
#> 3 41004
#> 4 41006
#> 5 41008
#> 6 41009
#> url
#> 1 http://dods.ndbc.noaa.gov/thredds/catalog/data/cwind/41001/catalog.html
#> 2 http://dods.ndbc.noaa.gov/thredds/catalog/data/cwind/41002/catalog.html
#> 3 http://dods.ndbc.noaa.gov/thredds/catalog/data/cwind/41004/catalog.html
#> 4 http://dods.ndbc.noaa.gov/thredds/catalog/data/cwind/41006/catalog.html
#> 5 http://dods.ndbc.noaa.gov/thredds/catalog/data/cwind/41008/catalog.html
#> 6 http://dods.ndbc.noaa.gov/thredds/catalog/data/cwind/41009/catalog.html
Get buoy data
With buoy you can get data for a particular dataset, buoy id, year, and datatype.
Get data for a buoy, specifying year and datatype
buoy(dataset = 'cwind', buoyid = 41001, year = 2008, datatype = "cc")
#> Dimensions (rows/cols): [1585 X 5]
#> 2 variables: [wind_dir, wind_spd]
#>
#> time latitude longitude wind_dir wind_spd
#> 1 2008-05-28T16:00:00Z 34.704 -72.734 230 8.6
#> 2 2008-05-28T16:10:00Z 34.704 -72.734 230 8.7
#> 3 2008-05-28T16:20:00Z 34.704 -72.734 229 8.5
#> 4 2008-05-28T16:30:00Z 34.704 -72.734 231 8.8
#> 5 2008-05-28T16:40:00Z 34.704 -72.734 236 8.5
#> 6 2008-05-28T16:50:00Z 34.704 -72.734 235 8.9
#> 7 2008-05-28T17:00:00Z 34.704 -72.734 233 8.2
#> 8 2008-05-28T17:10:00Z 34.704 -72.734 233 8.2
#> 9 2008-05-28T17:20:00Z 34.704 -72.734 231 8.3
#> 10 2008-05-28T17:30:00Z 34.704 -72.734 232 7.8
#> .. ... ... ... ... ...
More data
There are more NOAA data sources in noaa. Check out the various vignettes in the package.
Citing
To cite rnoaa in publications use:
Scott Chamberlain, Hart Edmund, and Karthik Ram (2015). rnoaa: NOAA climate data from R. R package version 0.4.2. https://github.com/ropensci/rnoaa
License and bugs
- License: MIT
- Report bugs at our Github repo for rnoaa